color spine width n_males weight
1 medium bad 28.3 8 3050
2 dark bad 22.5 0 1550
3 light good 26.0 9 2300
4 dark bad 24.8 0 2100
5 dark bad 26.0 4 2600
6 medium bad 23.8 0 2100
7 light good 26.5 0 2350
8 dark middle 24.7 0 1900Main Statistical Inquiries
Let us suppose we want to assess the following:
- Whether
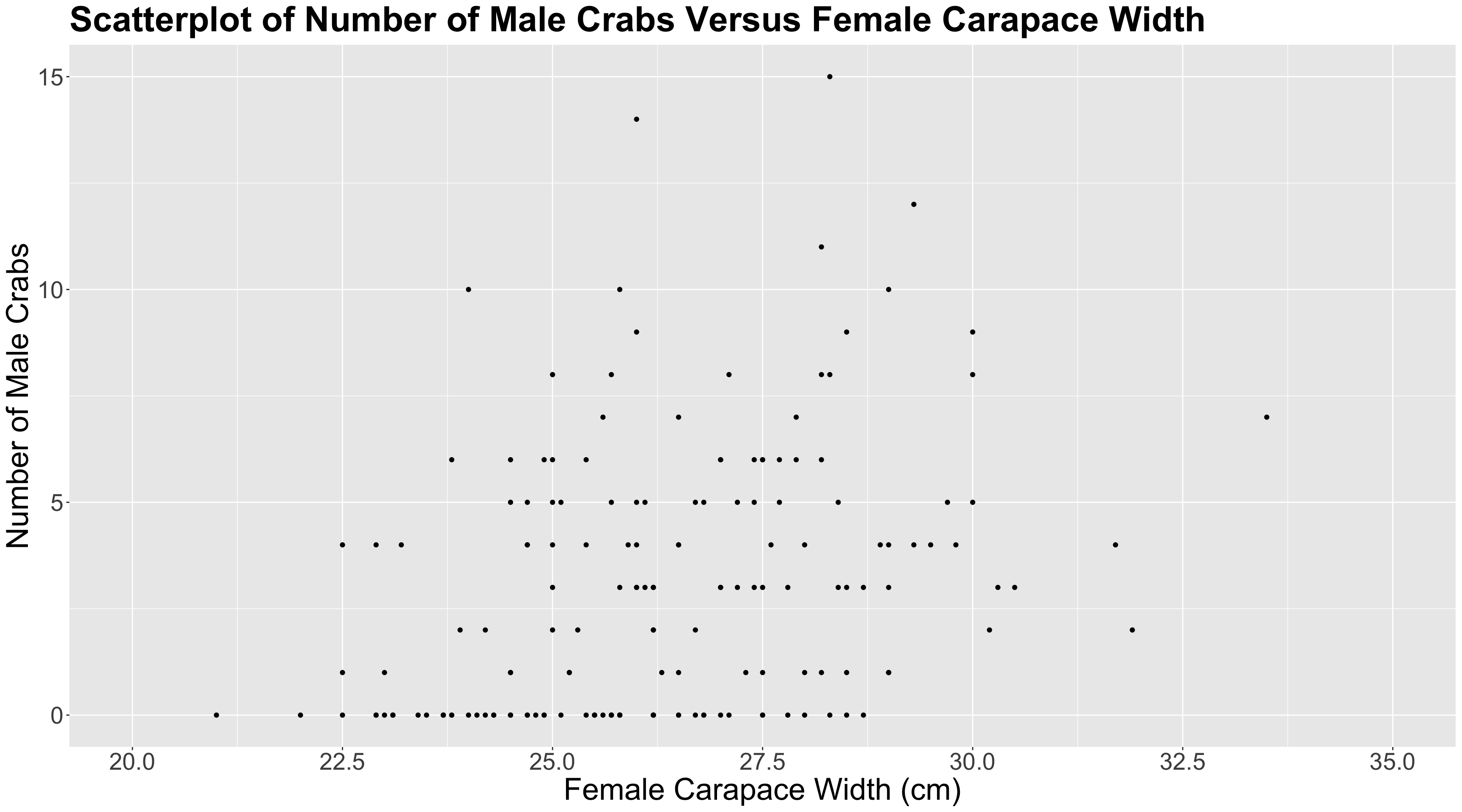
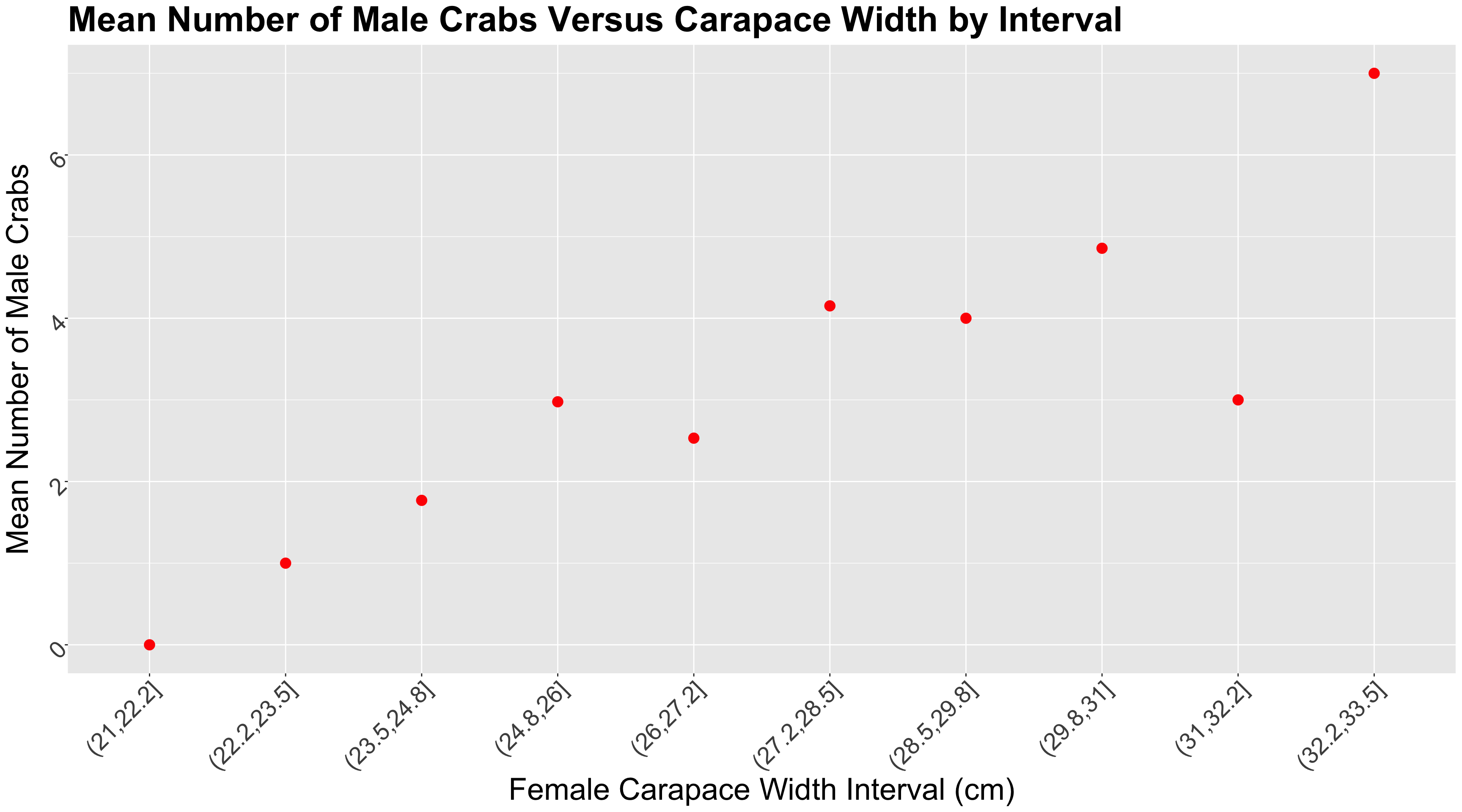
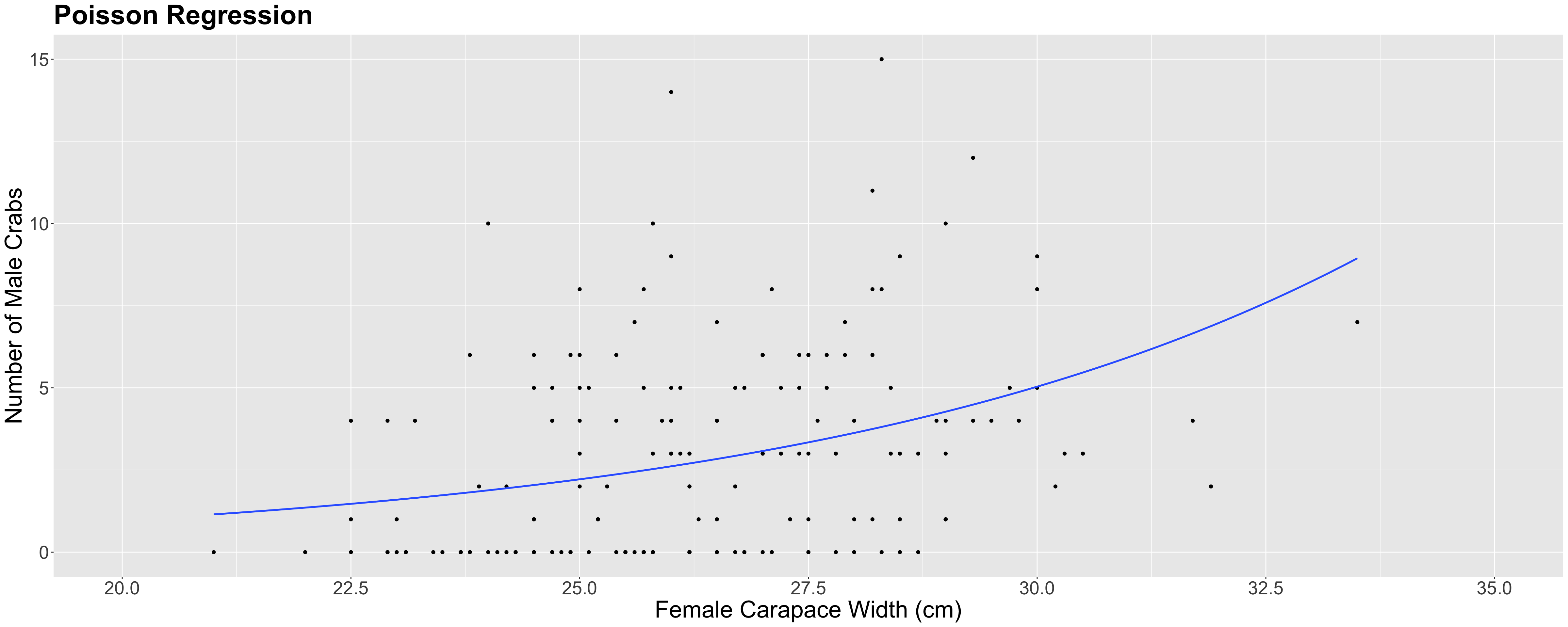
n_malesand female carapacewidthare statistically associated and by how much. - Whether
n_malesand femalecolorof the prosoma are statistically associated and by how much.