Main Statistical Inquiries
- We are interested in the following:
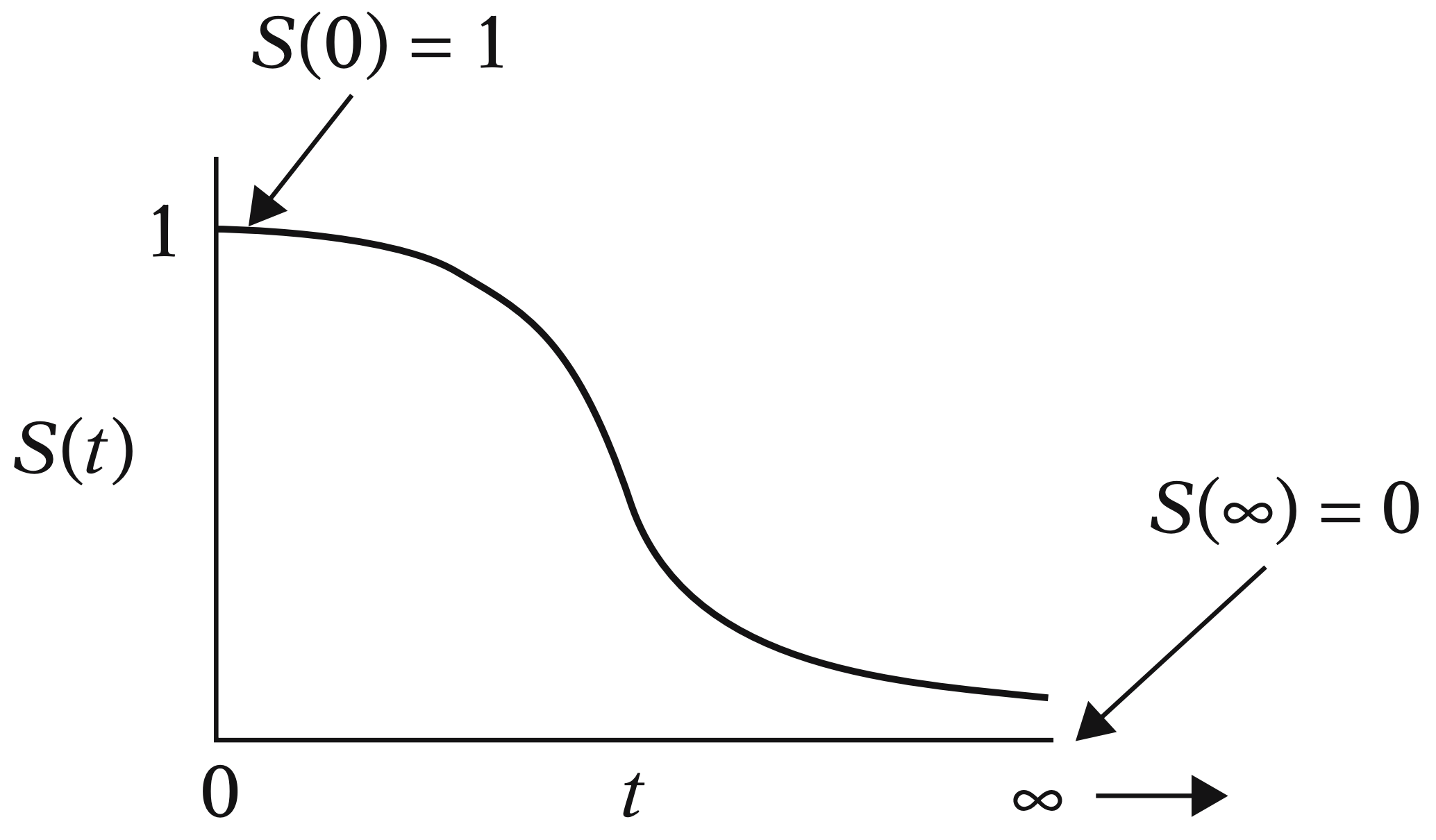
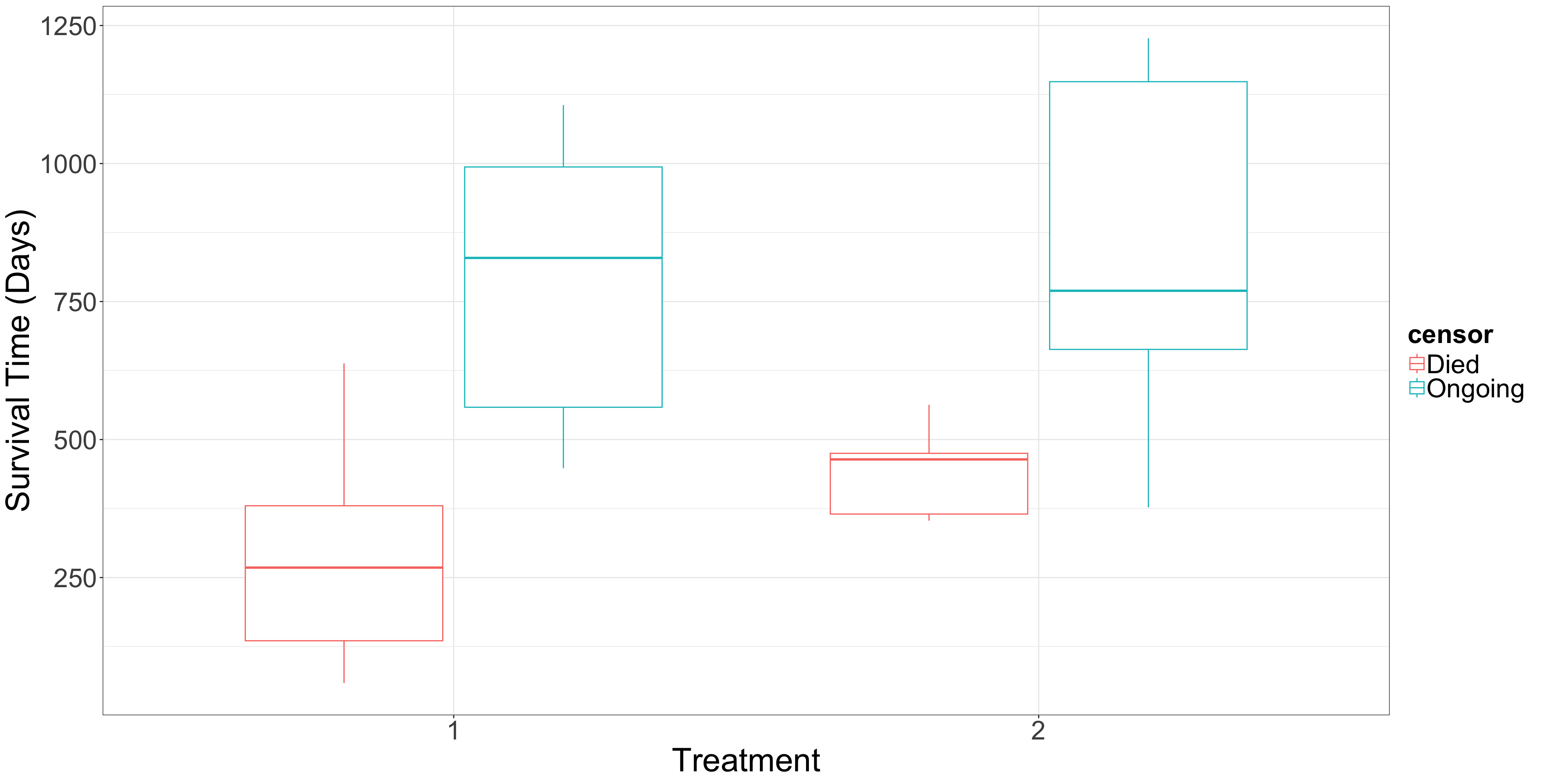
- Regardless of the cancer treatment used, is it possible to estimate the overall survival function for this population of cancer patients?
- Now, in terms of the specific cancer treatments, is there a statistical difference between both treatments? If so, can we quantify it? How would this be possible? Moreover, can we obtain the corresponding estimated survival functions per cancer treatment?